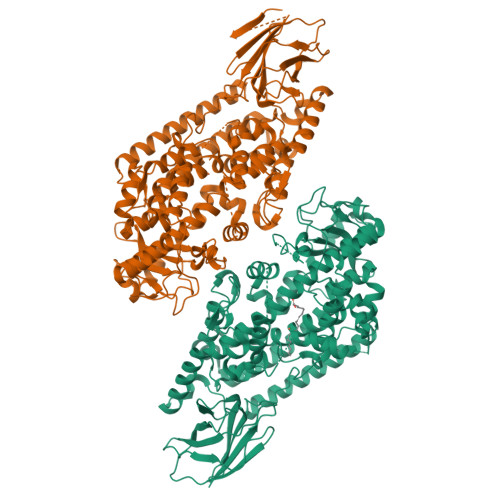
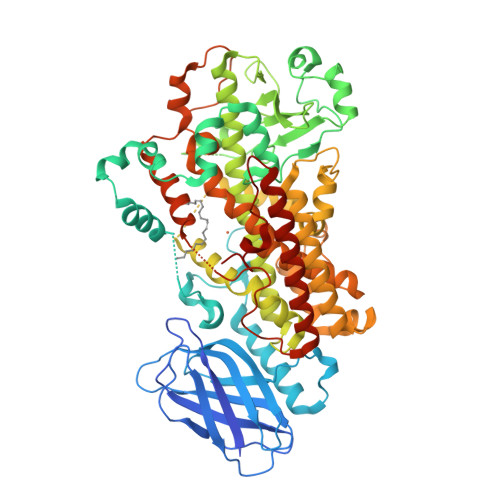
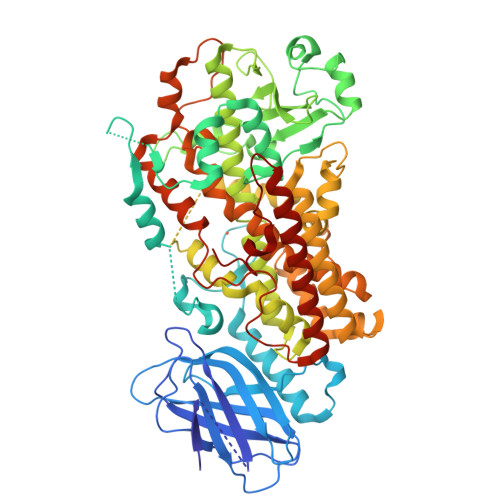
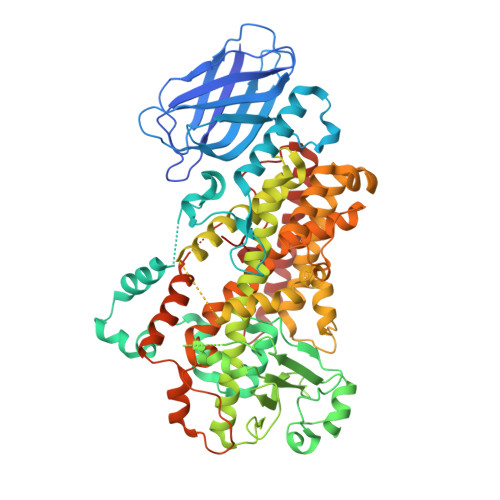
Conversion of human 5-lipoxygenase to a 15-lipoxygenase by a point mutation to mimic phosphorylation at Serine-663.
Gilbert, N.C., Rui, Z., Neau, D.B., Waight, M.T., Bartlett, S.G., Boeglin, W.E., Brash, A.R., Newcomer, M.E.(2012) FASEB J 26: 3222-3229
- PubMed: 22516296
- DOI: https://doi.org/10.1096/fj.12-205286
- Primary Citation of Related Structures:
3V92, 3V98, 3V99 - PubMed Abstract:
The enzyme 5-lipoxygenase (5-LOX) initiates biosynthesis of the proinflammatory leukotriene lipid mediators and, together with 15-LOX, is also required for synthesis of the anti-inflammatory lipoxins. The catalytic activity of 5-LOX is regulated through multiple mechanisms, including Ca(2+)-targeted membrane binding and phosphorylation at specific serine residues. To investigate the consequences of phosphorylation at S663, we mutated the residue to the phosphorylation mimic Asp, providing a homogenous preparation suitable for catalytic and structural studies. The S663D enzyme exhibits robust 15-LOX activity, as determined by spectrophotometric and HPLC analyses, with only traces of 5-LOX activity remaining; synthesis of the anti-inflammatory lipoxin A(4) from arachidonic acid is also detected. The crystal structure of the S663D mutant in the absence and presence of arachidonic acid (in the context of the previously reported Stable-5-LOX) reveals substantial remodeling of helices that define the active site so that the once fully encapsulated catalytic machinery is solvent accessible. Our results suggest that phosphorylation of 5-LOX at S663 could not only down-regulate leukotriene synthesis but also stimulate lipoxin production in inflammatory cells that do not express 15-LOX, thus redirecting lipid mediator biosynthesis to the production of proresolving mediators of inflammation.
Organizational Affiliation:
Department of Biological Sciences, Louisiana State University, Baton Rouge, LA 70803, USA.